Improved 16S Analysis on the One Codex Platform
Whether surveying the human microbiome, environmental sites, or other microbial communities, researchers have generally adopted one of two approaches – sequencing all of the DNA in a sample (WGS) or focusing on a specific marker gene (such as the 16S rDNA locus). While I tend to advocate for analyzing total DNA (via WGS), 16S sequencing remains popular for large sequencing projects due to its cost effectiveness.
Today, I’m excited to announce improved support for 16S analysis on the One Codex platform – which we hope will both make high quality 16S analysis more accessible, while also making it easier for researchers to integrate sample data from both 16S and shotgun sequencing projects.
Improved support for 16S and other markers on One Codex
Our new Marker Gene Database is specifically designed for marker gene sequencing and built using the most commonly used genes for microbial surveys, including 16S, ITS, 18S, and others.1 Here we are extending our k-mer-based exact alignment to marker genes, as has been demonstrated to good effect by other groups. The Marker Gene Database contains >300,000 curated gene records spanning the known microbial world – bacteria, archaea, fungi, protists, algae, etc. It builds on resources such as the NCBI Target Loci Project, as well as manually and automatically curated sequences from the broader NCBI nucleotide collection.
In building this marker database, we’ve sought to strike a balance between including the broadest possible number of known organisms while being careful to avoid contamination, mis-annotation, and other issues that plague the analysis of NGS data. You can see an example analysis using this new database here, which shows the results of this balanced curatorial effort.
Benchmarking the Marker Gene Database with a mock community
In order to assess the performance of the new Marker Gene Database, we analyzed a mock community of 59 strains of bacteria and archaea spanning 47 genera developed in a prior study (link). Mock communities are particularly useful for benchmarking because we know which organisms are truly present in the sample, and so every organism that a method detects can be classified as either a true positive or a false positive. While the authors constructed this mock community to more fully characterize the errors introduced by different library preparation techniques and sequencing instruments, here we’re interested in better understanding biases introduced in the downstream bioinformatics.
The below figure summarizes this analysis, and includes results for the new Marker Gene Database, our (whole genome only) One Codex Database, and the results reported by the study’s authors using the OTU-based tools used by QIIME, a commonly-used pipeline for 16S analysis.
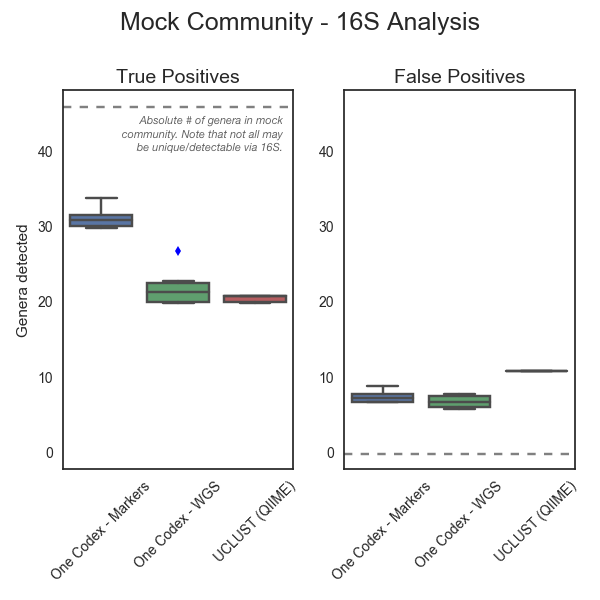
The Marker Gene Database (“One Codex – Markers”) had the highest rate of true positives (left panel),2 while UCLUST (the OTU tool used by QIIME) returned the highest number of false positives (right panel). We generally expect the One Codex Database of whole microbial genomes to perform well for marker gene data, because marker genes are contained within the genomes in the database. That was indeed the case here, with the One Codex Database of whole microbial genomes (“One Codex – WGS”) outperforming UCLUST with a lower rate of false positives. Overall, the Marker Gene Database gave the most accurate results, with the highest rate of true positives and the lowest rate of false positives.
Species-level detection via 16S
One of the biggest challenges of 16S analysis has been identifying bacteria down to the species level, not just the genus. We compared how accurately individual sequences could be assigned to each level of the taxonomy (phylum through species using One Codex, as well as three commonly-used tools for 16S analysis (RDP Classifier, Mothur-Greengenes, Mothur-SILVA)3.
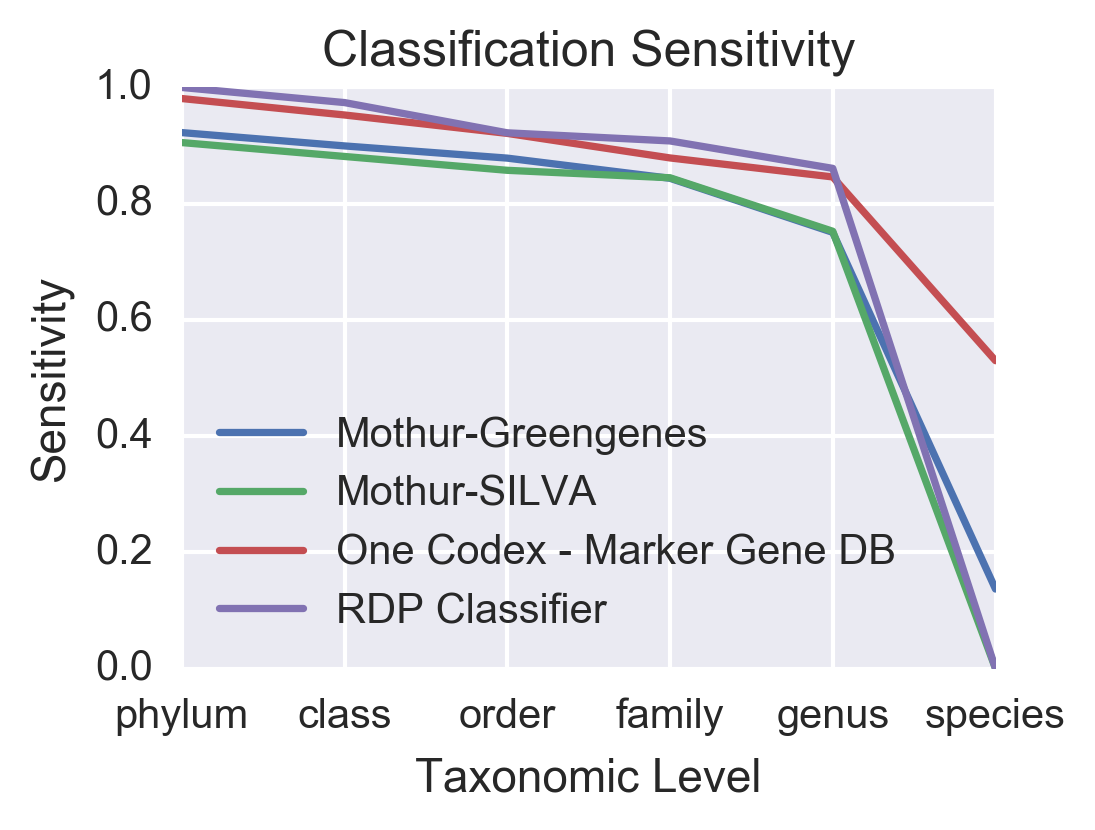
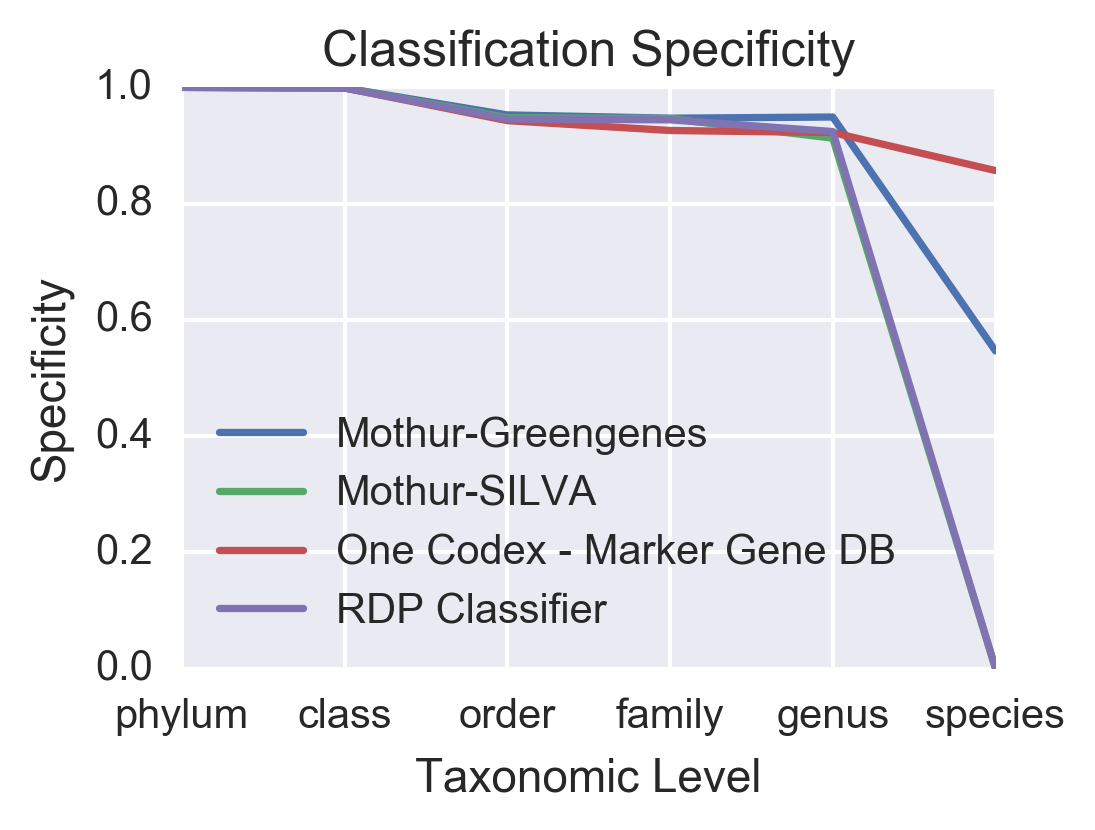 While the One Codex Marker Gene Database performs just as well as the other methods at assigning sequences down to the genus-level, it does a much better job of assigning sequences to the species level. While Mothur and the RDP Classifier assign less than 19% of sequences to the species-level, One Codex assigns over 52%, with specificity values approaching over 85%.
While the One Codex Marker Gene Database performs just as well as the other methods at assigning sequences down to the genus-level, it does a much better job of assigning sequences to the species level. While Mothur and the RDP Classifier assign less than 19% of sequences to the species-level, One Codex assigns over 52%, with specificity values approaching over 85%.
We think that some of the most important biology of the microbiome is driven by the different species that are present, and so I’m very gratified that we are able to provide species-level results for 16S data with a high level of accuracy.
(TODO: INSERT SECTION ON HOW TO KICK OFF 16S ANALYSES, HOW TO COMPARE RESULTS BETWEEN 16S AND SHOTGUN DATA, E.G, IN COMPARISON VIEW OR VIA API)
Questions? Comments?
As always, please feel free to drop me a note if you have any questions, feedback, or would like to discuss a project.
-- Sam Minot, Ph.D.
116S rDNA, 18S, 23S, 28S, 5S, ITS (Internal Transcribed Spacer), and gyrB (DNA gyrase subunit B). 2 Analysis was performed on the One Codex Platform, using an abundance threshold of 0.1% to account for sequencing error on the Illumina MiSeq. 3 RDP Classifier was run with version 2.11 (16S rRNA training set 15), Mothur version 1.37.5 run with default settings and minimum score 80.