Targeted Loci Post (long)
People that study the microbiome generally use two different genomic methods to analyze samples – sequencing all of the DNA in a sample (WGS) or targeting a specific marker gene (e.g. 16S rDNA). While WGS provides a high-resolution taxonomic and functional characterization of microbiome samples, 16S sequencing is a cost effective technique for broad community surveys across large collections of samples. Today, I’m excited to announce that One Codex is launching a powerful tool for 16S analysis – which we hope will make high-quality 16S analysis more accessible to a broad range of researchers.
Our approach to 16S – enabling robust, scalable, and portable analysis
The tools that analyze 16S data take either a “closed reference” approach (where sequences are compared to a fixed reference database of 16S sequences from known organisms), a “de novo” approach (where individual sequences are clustered by their pairwise similarity), or an “open reference” approach (a combination of the two). It’s important to note that for de novo and open reference analysis, adding new samples to a dataset generally requires the entire set of samples to be re-analyzed as a batch, which can be computationally intensive. We chose to take a closed reference approach for two reasons: 1) it tends to be more robust to changes in sampling depth and analysis parameters (e.g. the percent identity threshold used to create OTUs); and 2) comparing every sample against a fixed reference database allows us to perform cross-comparison across arbitrarily large number of samples, without having to downsample and recompute OTUs with every new dataset. We believe that this approach will enable researchers to easily analyze and compare across many thousands of samples, rapidly incorporating new datasets within a reproducible and deterministic analysis platform.
The One Codex Targeted Loci Database
The One Codex Targeted Loci Database is specifically designed for marker gene sequencing and built using the most commonly used genes for microbial surveys, including 16S, ITS, 18S, and others.1 The Targeted Loci Database contains ~250,000 curated gene records spanning the known microbial world – bacteria, archaea, fungi, protists, algae, etc. It builds on resources such as the NCBI Target Loci Project, as well as manually and automatically curated sequences from the broader NCBI nucleotide collection. We’ve sought to strike a balance between including the broadest possible number of known organisms while being careful to avoid contamination, mis-annotation, and other issues that can confound microbiome analysis2-4.
To analyze samples against the Targeted Loci Database, we sensitively align every read (using SNAP) and identify the best overall matches to the database, assigning each read to the most specific taxonomic grouping that the data supports (down to the species-level where appropriate). We then perform straightforward abundance-based filtering to minimize the number of false positive assignments that may be introduced by sequencing error.
Increased accuracy with the Targeted Loci Database
In order to measure the performance of the Targeted Loci Database, we analyzed a mock community constructed by Bokulich, et al.5 and analyzed in depth by Kopylova, et al.6. Mock communities are particularly useful for benchmarking because we know which organisms are truly present in the sample, and so every organism detected is either a true positive or a false positive. Moreover, the authors6 compared results from a wide range of 16S analysis tools, which we use here as a point of comparison.
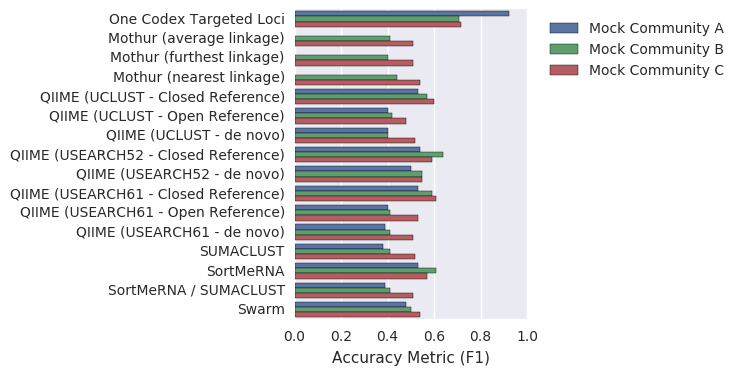
Looking across all of the analysis methods (using data from Table 2 of Kopylova, et al.6) the Targeted Loci Database (“One Codex”) performs quite well and has the highest overall accuracy (as measured by genus-level F score, which combines sensitivity and specificity) for all three of the test datasets. While all methods were able to detect the majority of organisms present, the One Codex Targeted Loci analysis generally reported the lowest number of false positives, providing the most accurate picture of the organisms present in these samples.
Robustly measuring community diversity
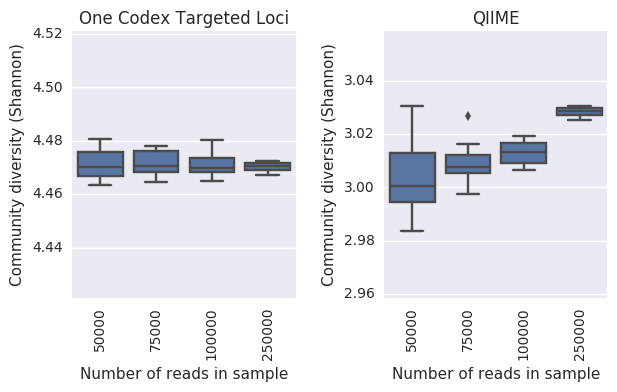
One of the most common uses of 16S analysis is measuring the overall taxonomic diversity of a microbiome sample or microbial community. This is the type of analysis that, for example, demonstrates that IBD is associated with reduced diversity in the gut microbiome7. We used a random subsampling approach to see how robustly the One Codex Targeted Loci analysis was able to measure community diversity at different levels of sequencing depth, compared to the commonly-used tool QIIME8. Across a group of 20 replicates each at 50K, 75K, 100K, and 250K reads per sample, the One Codex Targeted Loci analysis (left) provided an estimate of community diversity that was more consistent between replicates, relative to the output of QIIME (right). Moreover, the community diversity reported by QIIME increased as more reads were added to a sample, while the output of One Codex was not sensitive to sequencing depth.
Getting down to the species with 16S
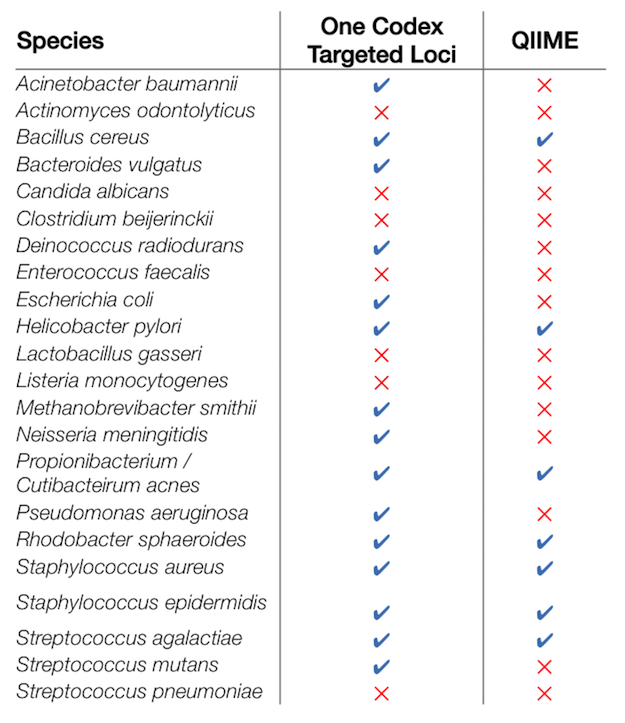
While 16S analysis is often performed at the genus-level, it is vastly preferable to know which species that are present in a community whenever possible. Although some species cannot be distinguished by 16S, we believe that a well-curated database and a sensitive detection algorithm will provide users with a greater ability to perform species-level detection. Looking at the species present in Mock Community A (table below), we found that the One Codex Targeted Loci analysis was able to detect 15 out of the 22 total, while QIIME8 only detected 7. While marker gene analysis doesn’t always contain species-level information, we believe that the One Codex Targeted Loci analysis does a good job of identifying those species that do have a distinct 16S gene.
Using the Targeted Loci Database
To run the Targeted Loci Database on your One Codex samples, go to the Run Analysis page, select your samples of interest using the menu on the left, and then click the Run button for the Targeted Loci Database on the right. This analysis runs for no additional cost with all samples uploaded to the One Codex platform.
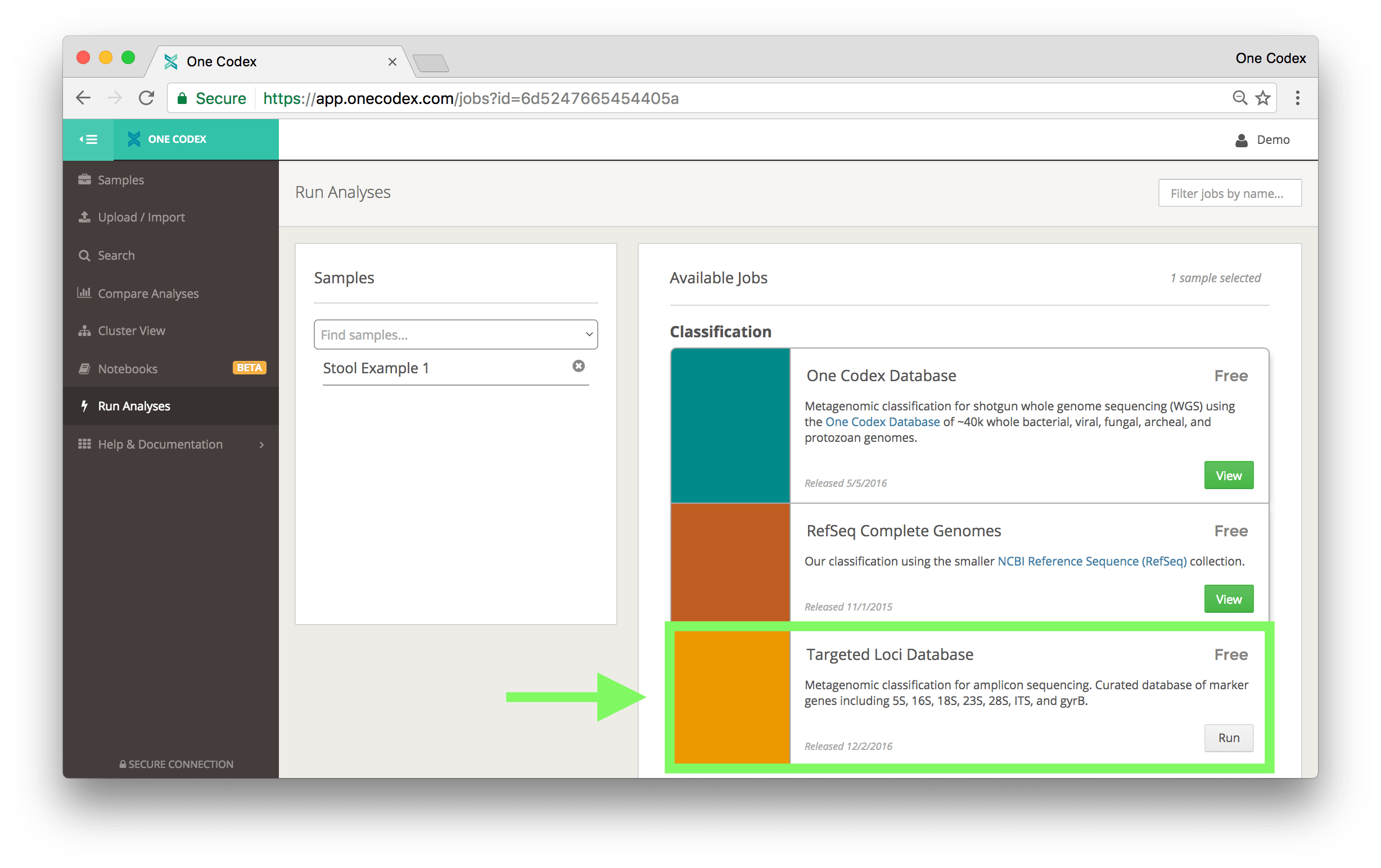
Finally, because this database uses the same NCBI taxonomy used by the One Codex Database, you can directly compare the results of WGS samples against 16S samples using the Compare Analysis Tool on the One Codex platform.
Questions? Comments?
As always, please feel free to drop me a note if you have any questions, feedback, or would like to discuss a project.
-- Sam Minot, Ph.D.
116S rDNA, 18S, 23S, 28S, 5S, ITS (Internal Transcribed Spacer), rpoB, and gyrB (DNA gyrase subunit B).
2 Salter SJ, et al. BMC Biol. 2014, 12:87. DOI: 10.1186/s12915-014-0087-z
3 Lusk RW. PLoS One. 2014, 9(10):e110808. doi: 10.1371/journal.pone.0110808
4 Merchant S, et al. PeerJ. 2014, 2:e675. doi: 10.7717/peerj.675
5 Bokulich NA, et al. Nature Methods 2014, 10(1) 57-59; DOI: 10.1038/nmeth.2276
6 Kopylova E, et al. mSystems 2016, 1(1) e00003-15; DOI: 10.1128/mSystems.00003-15
7 Kostic AD, et al. Gastroenterology 2014, 146(6); DOI: 10.1053/j.gastro.2014.02.009
8 QIIME (pick_open_reference_otus.py v1.9.1) was run with default settings, which include uclust for clustering and gg_13_8_otus/rep_set/97_otus.fasta as the default reference database.